diff options
| author | CoprDistGit <infra@openeuler.org> | 2023-06-09 08:10:21 +0000 |
|---|---|---|
| committer | CoprDistGit <infra@openeuler.org> | 2023-06-09 08:10:21 +0000 |
| commit | fde3e2c1fd6c0538b0be7004d823b2e25237353b (patch) | |
| tree | c68b7767a8cc68f48e4aa41b1d12489008f40fa6 | |
| parent | 696bad2568fd56a7ee7751a2670fa98cea6f0db3 (diff) | |
automatic import of python-immunemlopeneuler20.03
| -rw-r--r-- | .gitignore | 1 | ||||
| -rw-r--r-- | python-immuneml.spec | 713 | ||||
| -rw-r--r-- | sources | 1 |
3 files changed, 715 insertions, 0 deletions
@@ -0,0 +1 @@ +/immuneML-2.2.5.tar.gz diff --git a/python-immuneml.spec b/python-immuneml.spec new file mode 100644 index 0000000..6c6acdb --- /dev/null +++ b/python-immuneml.spec @@ -0,0 +1,713 @@ +%global _empty_manifest_terminate_build 0 +Name: python-immuneML +Version: 2.2.5 +Release: 1 +Summary: immuneML is a software platform for machine learning analysis of immune receptor repertoires. +License: GNU Affero General Public License v3 +URL: https://github.com/uio-bmi/immuneML +Source0: https://mirrors.aliyun.com/pypi/web/packages/78/0a/0f6dd04d3ac651f2b7d1669b486940f0ce5066612f8177642a35839bdb0a/immuneML-2.2.5.tar.gz +BuildArch: noarch + +Requires: python3-numpy +Requires: python3-pytest +Requires: python3-pandas +Requires: python3-PyYAML +Requires: python3-scikit-learn +Requires: python3-gensim +Requires: python3-matplotlib +Requires: python3-editdistance +Requires: python3-regex +Requires: python3-tzlocal +Requires: python3-airr +Requires: python3-fishersapi +Requires: python3-pystache +Requires: python3-torch +Requires: python3-dill +Requires: python3-tensorboard +Requires: python3-plotly +Requires: python3-logomaker +Requires: python3-matplotlib-venn +Requires: python3-scipy +Requires: python3-Cython +Requires: python3-parasail +Requires: python3-tcrdist3 + +%description +# immuneML + + + +[](https://docs.airr-community.org/en/stable/swtools/airr_swtools_standard.html) + + +immuneML is a platform for machine learning-based analysis and +classification of adaptive immune receptors and repertoires (AIRR). + +It supports the analyses of experimental B- and T-cell receptor data, +as well as synthetic data for benchmarking purposes. + +In immuneML, users can define flexible workflows supporting different +machine learning libraries (such as scikit-learn or PyTorch), benchmarking of different approaches, numerous reports +of data characteristics, ML algorithms and their predictions, and +visualizations of results. + +Additionally, users can extend the platform by defining their own data +representations, ML models, reports and visualizations. + + +Useful links: +- Main website: https://immuneml.uio.no +- Documentation: https://docs.immuneml.uio.no +- Galaxy web interface: https://galaxy.immuneml.uiocloud.no + + + +## Installation + +immuneML can be installed directly [using pip](<https://pypi.org/project/immuneML/>). +immuneML uses Python 3.7 or 3.8, we recommend installing immuneML inside a virtual environment +with one of these Python versions. + +For more detailed instructions (virtual environment, troubleshooting, Docker, developer installation), please see the [installation documentation](https://docs.immuneml.uio.no/installation/install_with_package_manager.html). + +### Installation using pip + + +To install the immuneML core package, run: + +```bash +pip install immuneML +``` + +Alternatively, to use the TCRdistClassifier ML method and corresponding TCRdistMotifDiscovery report, install immuneML with the optional TCRdist extra: + +```bash +pip install immuneML[TCRdist] +``` + +Optionally, if you want to use the DeepRC ML method and and corresponding DeepRCMotifDiscovery report, you also +have to install DeepRC dependencies using the [requirements_DeepRC.txt](https://raw.githubusercontent.com/uio-bmi/immuneML/master/requirements_DeepRC.txt) file. +Important note: DeepRC uses PyTorch functionalities that depend on GPU. Therefore, DeepRC does not work on a CPU. +To install the DeepRC dependencies, run: + +```bash +pip install -r requirements_DeepRC.txt --no-dependencies +``` + +### Validating the installation + +To validate the installation, run: + +```bash +immune-ml -h +``` + +This should display a help message explaining immuneML usage. + +To quickly test out whether immuneML is able to run, try running the quickstart command: + +```bash +immune-ml-quickstart ./quickstart_results/ +``` + +This will generate a synthetic dataset and run a simple machine machine learning analysis +on the generated data. The results folder will contain two sub-folders: one for the generated dataset (`synthetic_dataset`) +and one for the results of the machine learning analysis (`machine_learning_analysis`). +The files named `specs.yaml` are the input files for immuneML that describe how to generate +the dataset and how to do the machine learning analysis. The `index.html` files can be used +to navigate through all the results that were produced. + +## Usage + +### Quickstart + +The quickest way to familiarize yourself with immuneML usage is to follow +one of the [Quickstart tutorials](https://docs.immuneml.uio.no/quickstart.html). +These tutorials provide a step-by-step guide on how to use immuneML for a +simple machine learning analysis on an adaptive immune receptor repertoire (AIRR) dataset, +using either the command line tool or the [Galaxy web interface](https://galaxy.immuneml.uiocloud.no). + + +### Overview of input, analyses and results + +The figure below shows an overview of immuneML usage. +All parameters for an immuneML analysis are defined in the a YAML specification file. +In this file, the settings of the analysis components are defined (also known as `definitions`, +shown in six different colors in the figure). +Additionally, the YAML file describes one or more `instructions`, which are workflows that are +applied to the defined analysis components. +Each instruction uses at least a dataset component, and optionally additional components. +AIRR datasets may either be [imported from files](https://docs.immuneml.uio.no/tutorials/how_to_import_the_data_to_immuneML.html), +or [generated synthetically](https://docs.immuneml.uio.no/tutorials/how_to_generate_a_random_repertoire_dataset.html) during runtime. + +Each instruction produces different types of results, including trained ML models, +ML model predictions on a given dataset, plots or other reports describing the +dataset or trained models, and modified datasets. +To navigate over the results, immuneML generates a summary HTML file. + + +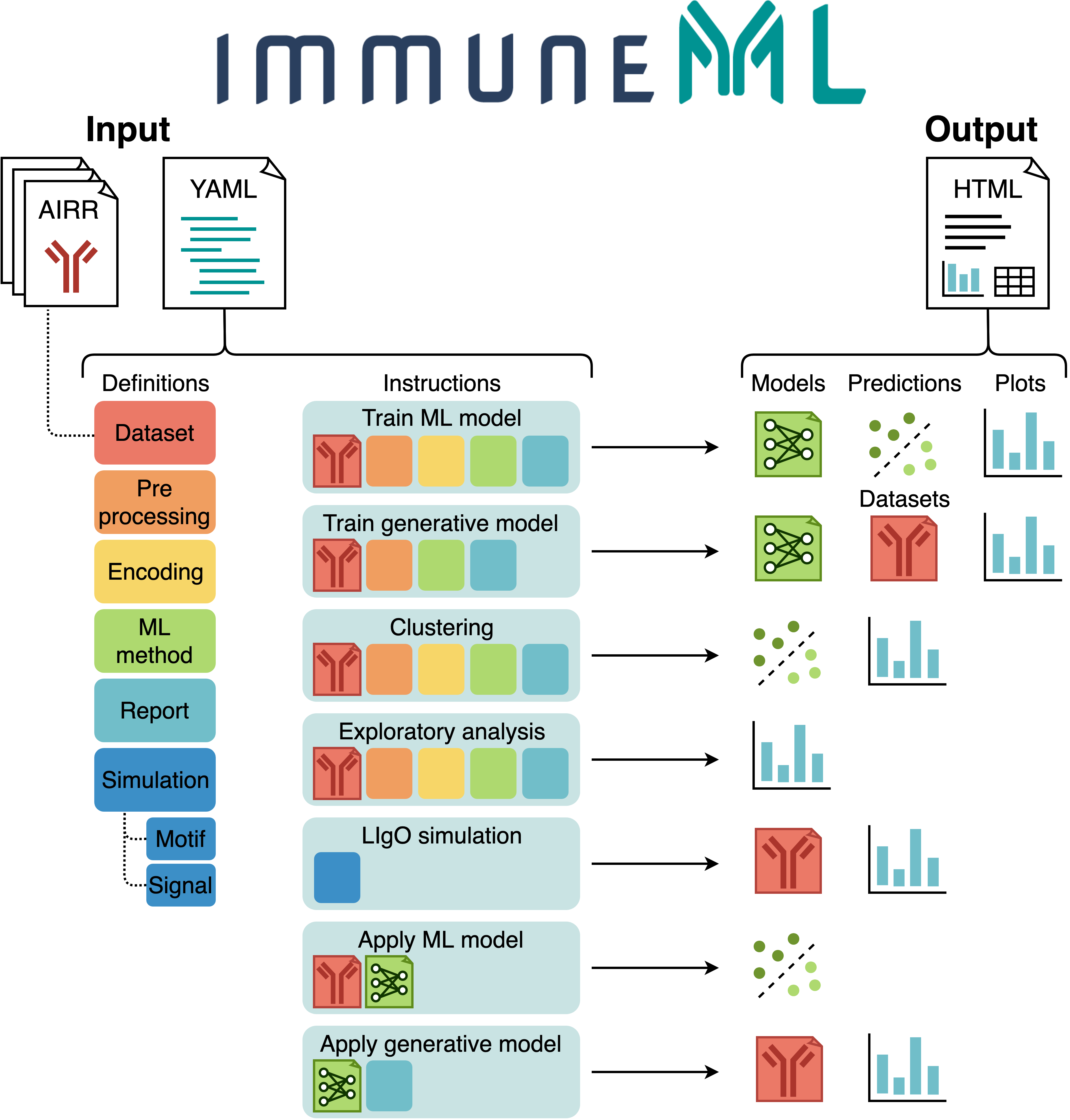 + +For a detailed explanation of the YAML specification file, see the tutorial [How to specify an analysis with YAML](https://docs.immuneml.uio.no/tutorials/how_to_specify_an_analysis_with_yaml.html). + +See also the following tutorials for specific instructions: +- [Training ML models](https://docs.immuneml.uio.no/tutorials/how_to_train_and_assess_a_receptor_or_repertoire_classifier.html) for repertoire classification (e.g., disease prediction) or receptor sequence classification (e.g., antigen binding prediction). In immuneML, the performance of different machine learning (ML) settings can be compared by nested cross-validation. These ML settings consist of data preprocessing steps, encodings and ML models and their hyperparameters. +- [Exploratory analysis](https://docs.immuneml.uio.no/tutorials/how_to_perform_exploratory_analysis.html) of datasets by applying preprocessing and encoding, and plotting descriptive statistics without training ML models. +- [Simulating](https://docs.immuneml.uio.no/tutorials/how_to_simulate_antigen_signals_in_airr_datasets.html) immune events, such as disease states, into experimental or synthetic repertoire datasets. By implanting known immune signals into a given dataset, a ground truth benchmarking dataset is created. Such a dataset can be used to test the performance of ML settings under known conditions. +- [Applying trained ML models](https://docs.immuneml.uio.no/tutorials/how_to_apply_to_new_data.html) to new datasets with unknown class labels. +- And [other tutorials](https://docs.immuneml.uio.no/tutorials.html) + + +### Command line usage + +The `immune-ml` command takes only two parameters: the YAML specification file and a result path. +An example is given here: + +```bash +immune-ml path/to/specification.yaml result/folder/path/ +``` + +For each instruction specified in the YAML specification file, a subfolder is created in the +`result/folder/path`. Each subfolder will contain: +- An `index.html` file which shows an overview of the results produced by that instruction. Inspecting the results of an immuneML analysis typically starts here. +- A copy of the used YAML specification (`full_specification.yaml`) with all default parameters explicitly set. +- A folder containing all raw results produced by the instruction. +- A folder containing the imported dataset(s) in optimized binary (Pickle) format. + +## Support + +We will prioritize fixing important bugs, and try to answer any questions as soon as possible. We may implement suggested features and enhancements as time permits. + +If you run into problems when using immuneML, please see [the documentation](https://docs.immuneml.uio.no/latest/). In particular, we recommend you check out: +- The [Quickstart tutorial](https://docs.immuneml.uio.no/latest/quickstart.html) for new users +- The [Troubleshooting](https://docs.immuneml.uio.no/latest/troubleshooting.html) page + +If this does not answer your question, you can contact us via: +- Twitter [`@immuneml`](https://twitter.com/immuneml) +- Email [`contact@immuneml.uio.no`](mailto:contact@immuneml.uio.no) + +To report a potential bug or suggest new features, please [submit an issue on GitHub](https://github.com/uio-bmi/immuneML/issues). + +If you would like to make contributions, for example by adding a new ML method, encoding, report or preprocessing, please [see our developer documentation](https://docs.immuneml.uio.no/latest/developer_docs.html) and [submit a pull request](https://github.com/uio-bmi/compairr/pulls). + +## Requirements + +- [Python 3.7 or 3.8](https://www.python.org/) +- Python packages: + - [airr](https://pypi.org/project/airr/) (1 or higher) + - [dill](https://pypi.org/project/dill/) (0.3 or higher) + - [editdistance](https://pypi.org/project/editdistance/) (0.5.3 or higher) + - [fishersapi](https://pypi.org/project/fishersapi/) + - [gensim](https://pypi.org/project/gensim/) (3.8 or higher, < 4) + - [logomaker](https://pypi.org/project/logomaker/) (0.8 or higher) + - [matplotlib](https://matplotlib.org) (3.1 or higher) + - [matplotlib-venn](https://pypi.org/project/matplotlib-venn/) (0.11 or higher) + - [numpy](https://www.numpy.org/) (1.18 or higher, but at most 1.23.5) + - [pandas](https://pandas.pydata.org/) (1 or higher) + - [plotly](https://plotly.com/python/) (4 or higher) + - [pystache](https://pypi.org/project/pystache/) (0.5.4) + - [Pytorch](https://pytorch.org/) (1.5.1 or higher) + - [PyYAML](https://pyyaml.org) (5.3 or higher) + - [regex](https://pypi.org/project/regex/) + - [scikit-learn](https://scikit-learn.org/) (0.23 or higher) + - [scipy](https://www.scipy.org) + - [tensorboard](https://www.tensorflow.org/tensorboard) (1.14.0 or higher) + - [tzlocal](https://pypi.org/project/tzlocal/) +- Optional dependencies when using DeepRC: + - [DeepRC](https://github.com/ml-jku/DeepRC) (0.0.1) + - [widis-lstm-tools](https://github.com/widmi/widis-lstm-tools) (0.4) + - [tqdm](https://tqdm.github.io/) (0.24 or higher) + - [h5py](https://www.h5py.org/) (2.10.0 or lower when using DeepRC 0.0.1) +- Optional dependencies when using TCRdist: + - [parasail](https://pypi.org/project/parasail/) (1.2) + - [tcrdist3](https://github.com/kmayerb/tcrdist3) (0.1.6 or higher) + +# Citing immuneML + +If you are using immuneML in any published work, please cite: + +Pavlović, M., Scheffer, L., Motwani, K. et al. The immuneML ecosystem for machine learning analysis of adaptive immune +receptor repertoires. Nat Mach Intell 3, 936–944 (2021). https://doi.org/10.1038/s42256-021-00413-z + + + +<hr> + + +© Copyright 2021-2022, Milena Pavlovic, Lonneke Scheffer, Keshav Motwani, Victor Greiff, Geir Kjetil Sandve + + + + + + +%package -n python3-immuneML +Summary: immuneML is a software platform for machine learning analysis of immune receptor repertoires. +Provides: python-immuneML +BuildRequires: python3-devel +BuildRequires: python3-setuptools +BuildRequires: python3-pip +%description -n python3-immuneML +# immuneML + + + +[](https://docs.airr-community.org/en/stable/swtools/airr_swtools_standard.html) + + +immuneML is a platform for machine learning-based analysis and +classification of adaptive immune receptors and repertoires (AIRR). + +It supports the analyses of experimental B- and T-cell receptor data, +as well as synthetic data for benchmarking purposes. + +In immuneML, users can define flexible workflows supporting different +machine learning libraries (such as scikit-learn or PyTorch), benchmarking of different approaches, numerous reports +of data characteristics, ML algorithms and their predictions, and +visualizations of results. + +Additionally, users can extend the platform by defining their own data +representations, ML models, reports and visualizations. + + +Useful links: +- Main website: https://immuneml.uio.no +- Documentation: https://docs.immuneml.uio.no +- Galaxy web interface: https://galaxy.immuneml.uiocloud.no + + + +## Installation + +immuneML can be installed directly [using pip](<https://pypi.org/project/immuneML/>). +immuneML uses Python 3.7 or 3.8, we recommend installing immuneML inside a virtual environment +with one of these Python versions. + +For more detailed instructions (virtual environment, troubleshooting, Docker, developer installation), please see the [installation documentation](https://docs.immuneml.uio.no/installation/install_with_package_manager.html). + +### Installation using pip + + +To install the immuneML core package, run: + +```bash +pip install immuneML +``` + +Alternatively, to use the TCRdistClassifier ML method and corresponding TCRdistMotifDiscovery report, install immuneML with the optional TCRdist extra: + +```bash +pip install immuneML[TCRdist] +``` + +Optionally, if you want to use the DeepRC ML method and and corresponding DeepRCMotifDiscovery report, you also +have to install DeepRC dependencies using the [requirements_DeepRC.txt](https://raw.githubusercontent.com/uio-bmi/immuneML/master/requirements_DeepRC.txt) file. +Important note: DeepRC uses PyTorch functionalities that depend on GPU. Therefore, DeepRC does not work on a CPU. +To install the DeepRC dependencies, run: + +```bash +pip install -r requirements_DeepRC.txt --no-dependencies +``` + +### Validating the installation + +To validate the installation, run: + +```bash +immune-ml -h +``` + +This should display a help message explaining immuneML usage. + +To quickly test out whether immuneML is able to run, try running the quickstart command: + +```bash +immune-ml-quickstart ./quickstart_results/ +``` + +This will generate a synthetic dataset and run a simple machine machine learning analysis +on the generated data. The results folder will contain two sub-folders: one for the generated dataset (`synthetic_dataset`) +and one for the results of the machine learning analysis (`machine_learning_analysis`). +The files named `specs.yaml` are the input files for immuneML that describe how to generate +the dataset and how to do the machine learning analysis. The `index.html` files can be used +to navigate through all the results that were produced. + +## Usage + +### Quickstart + +The quickest way to familiarize yourself with immuneML usage is to follow +one of the [Quickstart tutorials](https://docs.immuneml.uio.no/quickstart.html). +These tutorials provide a step-by-step guide on how to use immuneML for a +simple machine learning analysis on an adaptive immune receptor repertoire (AIRR) dataset, +using either the command line tool or the [Galaxy web interface](https://galaxy.immuneml.uiocloud.no). + + +### Overview of input, analyses and results + +The figure below shows an overview of immuneML usage. +All parameters for an immuneML analysis are defined in the a YAML specification file. +In this file, the settings of the analysis components are defined (also known as `definitions`, +shown in six different colors in the figure). +Additionally, the YAML file describes one or more `instructions`, which are workflows that are +applied to the defined analysis components. +Each instruction uses at least a dataset component, and optionally additional components. +AIRR datasets may either be [imported from files](https://docs.immuneml.uio.no/tutorials/how_to_import_the_data_to_immuneML.html), +or [generated synthetically](https://docs.immuneml.uio.no/tutorials/how_to_generate_a_random_repertoire_dataset.html) during runtime. + +Each instruction produces different types of results, including trained ML models, +ML model predictions on a given dataset, plots or other reports describing the +dataset or trained models, and modified datasets. +To navigate over the results, immuneML generates a summary HTML file. + + + + +For a detailed explanation of the YAML specification file, see the tutorial [How to specify an analysis with YAML](https://docs.immuneml.uio.no/tutorials/how_to_specify_an_analysis_with_yaml.html). + +See also the following tutorials for specific instructions: +- [Training ML models](https://docs.immuneml.uio.no/tutorials/how_to_train_and_assess_a_receptor_or_repertoire_classifier.html) for repertoire classification (e.g., disease prediction) or receptor sequence classification (e.g., antigen binding prediction). In immuneML, the performance of different machine learning (ML) settings can be compared by nested cross-validation. These ML settings consist of data preprocessing steps, encodings and ML models and their hyperparameters. +- [Exploratory analysis](https://docs.immuneml.uio.no/tutorials/how_to_perform_exploratory_analysis.html) of datasets by applying preprocessing and encoding, and plotting descriptive statistics without training ML models. +- [Simulating](https://docs.immuneml.uio.no/tutorials/how_to_simulate_antigen_signals_in_airr_datasets.html) immune events, such as disease states, into experimental or synthetic repertoire datasets. By implanting known immune signals into a given dataset, a ground truth benchmarking dataset is created. Such a dataset can be used to test the performance of ML settings under known conditions. +- [Applying trained ML models](https://docs.immuneml.uio.no/tutorials/how_to_apply_to_new_data.html) to new datasets with unknown class labels. +- And [other tutorials](https://docs.immuneml.uio.no/tutorials.html) + + +### Command line usage + +The `immune-ml` command takes only two parameters: the YAML specification file and a result path. +An example is given here: + +```bash +immune-ml path/to/specification.yaml result/folder/path/ +``` + +For each instruction specified in the YAML specification file, a subfolder is created in the +`result/folder/path`. Each subfolder will contain: +- An `index.html` file which shows an overview of the results produced by that instruction. Inspecting the results of an immuneML analysis typically starts here. +- A copy of the used YAML specification (`full_specification.yaml`) with all default parameters explicitly set. +- A folder containing all raw results produced by the instruction. +- A folder containing the imported dataset(s) in optimized binary (Pickle) format. + +## Support + +We will prioritize fixing important bugs, and try to answer any questions as soon as possible. We may implement suggested features and enhancements as time permits. + +If you run into problems when using immuneML, please see [the documentation](https://docs.immuneml.uio.no/latest/). In particular, we recommend you check out: +- The [Quickstart tutorial](https://docs.immuneml.uio.no/latest/quickstart.html) for new users +- The [Troubleshooting](https://docs.immuneml.uio.no/latest/troubleshooting.html) page + +If this does not answer your question, you can contact us via: +- Twitter [`@immuneml`](https://twitter.com/immuneml) +- Email [`contact@immuneml.uio.no`](mailto:contact@immuneml.uio.no) + +To report a potential bug or suggest new features, please [submit an issue on GitHub](https://github.com/uio-bmi/immuneML/issues). + +If you would like to make contributions, for example by adding a new ML method, encoding, report or preprocessing, please [see our developer documentation](https://docs.immuneml.uio.no/latest/developer_docs.html) and [submit a pull request](https://github.com/uio-bmi/compairr/pulls). + +## Requirements + +- [Python 3.7 or 3.8](https://www.python.org/) +- Python packages: + - [airr](https://pypi.org/project/airr/) (1 or higher) + - [dill](https://pypi.org/project/dill/) (0.3 or higher) + - [editdistance](https://pypi.org/project/editdistance/) (0.5.3 or higher) + - [fishersapi](https://pypi.org/project/fishersapi/) + - [gensim](https://pypi.org/project/gensim/) (3.8 or higher, < 4) + - [logomaker](https://pypi.org/project/logomaker/) (0.8 or higher) + - [matplotlib](https://matplotlib.org) (3.1 or higher) + - [matplotlib-venn](https://pypi.org/project/matplotlib-venn/) (0.11 or higher) + - [numpy](https://www.numpy.org/) (1.18 or higher, but at most 1.23.5) + - [pandas](https://pandas.pydata.org/) (1 or higher) + - [plotly](https://plotly.com/python/) (4 or higher) + - [pystache](https://pypi.org/project/pystache/) (0.5.4) + - [Pytorch](https://pytorch.org/) (1.5.1 or higher) + - [PyYAML](https://pyyaml.org) (5.3 or higher) + - [regex](https://pypi.org/project/regex/) + - [scikit-learn](https://scikit-learn.org/) (0.23 or higher) + - [scipy](https://www.scipy.org) + - [tensorboard](https://www.tensorflow.org/tensorboard) (1.14.0 or higher) + - [tzlocal](https://pypi.org/project/tzlocal/) +- Optional dependencies when using DeepRC: + - [DeepRC](https://github.com/ml-jku/DeepRC) (0.0.1) + - [widis-lstm-tools](https://github.com/widmi/widis-lstm-tools) (0.4) + - [tqdm](https://tqdm.github.io/) (0.24 or higher) + - [h5py](https://www.h5py.org/) (2.10.0 or lower when using DeepRC 0.0.1) +- Optional dependencies when using TCRdist: + - [parasail](https://pypi.org/project/parasail/) (1.2) + - [tcrdist3](https://github.com/kmayerb/tcrdist3) (0.1.6 or higher) + +# Citing immuneML + +If you are using immuneML in any published work, please cite: + +Pavlović, M., Scheffer, L., Motwani, K. et al. The immuneML ecosystem for machine learning analysis of adaptive immune +receptor repertoires. Nat Mach Intell 3, 936–944 (2021). https://doi.org/10.1038/s42256-021-00413-z + + + +<hr> + + +© Copyright 2021-2022, Milena Pavlovic, Lonneke Scheffer, Keshav Motwani, Victor Greiff, Geir Kjetil Sandve + + + + + + +%package help +Summary: Development documents and examples for immuneML +Provides: python3-immuneML-doc +%description help +# immuneML + + + +[](https://docs.airr-community.org/en/stable/swtools/airr_swtools_standard.html) + + +immuneML is a platform for machine learning-based analysis and +classification of adaptive immune receptors and repertoires (AIRR). + +It supports the analyses of experimental B- and T-cell receptor data, +as well as synthetic data for benchmarking purposes. + +In immuneML, users can define flexible workflows supporting different +machine learning libraries (such as scikit-learn or PyTorch), benchmarking of different approaches, numerous reports +of data characteristics, ML algorithms and their predictions, and +visualizations of results. + +Additionally, users can extend the platform by defining their own data +representations, ML models, reports and visualizations. + + +Useful links: +- Main website: https://immuneml.uio.no +- Documentation: https://docs.immuneml.uio.no +- Galaxy web interface: https://galaxy.immuneml.uiocloud.no + + + +## Installation + +immuneML can be installed directly [using pip](<https://pypi.org/project/immuneML/>). +immuneML uses Python 3.7 or 3.8, we recommend installing immuneML inside a virtual environment +with one of these Python versions. + +For more detailed instructions (virtual environment, troubleshooting, Docker, developer installation), please see the [installation documentation](https://docs.immuneml.uio.no/installation/install_with_package_manager.html). + +### Installation using pip + + +To install the immuneML core package, run: + +```bash +pip install immuneML +``` + +Alternatively, to use the TCRdistClassifier ML method and corresponding TCRdistMotifDiscovery report, install immuneML with the optional TCRdist extra: + +```bash +pip install immuneML[TCRdist] +``` + +Optionally, if you want to use the DeepRC ML method and and corresponding DeepRCMotifDiscovery report, you also +have to install DeepRC dependencies using the [requirements_DeepRC.txt](https://raw.githubusercontent.com/uio-bmi/immuneML/master/requirements_DeepRC.txt) file. +Important note: DeepRC uses PyTorch functionalities that depend on GPU. Therefore, DeepRC does not work on a CPU. +To install the DeepRC dependencies, run: + +```bash +pip install -r requirements_DeepRC.txt --no-dependencies +``` + +### Validating the installation + +To validate the installation, run: + +```bash +immune-ml -h +``` + +This should display a help message explaining immuneML usage. + +To quickly test out whether immuneML is able to run, try running the quickstart command: + +```bash +immune-ml-quickstart ./quickstart_results/ +``` + +This will generate a synthetic dataset and run a simple machine machine learning analysis +on the generated data. The results folder will contain two sub-folders: one for the generated dataset (`synthetic_dataset`) +and one for the results of the machine learning analysis (`machine_learning_analysis`). +The files named `specs.yaml` are the input files for immuneML that describe how to generate +the dataset and how to do the machine learning analysis. The `index.html` files can be used +to navigate through all the results that were produced. + +## Usage + +### Quickstart + +The quickest way to familiarize yourself with immuneML usage is to follow +one of the [Quickstart tutorials](https://docs.immuneml.uio.no/quickstart.html). +These tutorials provide a step-by-step guide on how to use immuneML for a +simple machine learning analysis on an adaptive immune receptor repertoire (AIRR) dataset, +using either the command line tool or the [Galaxy web interface](https://galaxy.immuneml.uiocloud.no). + + +### Overview of input, analyses and results + +The figure below shows an overview of immuneML usage. +All parameters for an immuneML analysis are defined in the a YAML specification file. +In this file, the settings of the analysis components are defined (also known as `definitions`, +shown in six different colors in the figure). +Additionally, the YAML file describes one or more `instructions`, which are workflows that are +applied to the defined analysis components. +Each instruction uses at least a dataset component, and optionally additional components. +AIRR datasets may either be [imported from files](https://docs.immuneml.uio.no/tutorials/how_to_import_the_data_to_immuneML.html), +or [generated synthetically](https://docs.immuneml.uio.no/tutorials/how_to_generate_a_random_repertoire_dataset.html) during runtime. + +Each instruction produces different types of results, including trained ML models, +ML model predictions on a given dataset, plots or other reports describing the +dataset or trained models, and modified datasets. +To navigate over the results, immuneML generates a summary HTML file. + + + + +For a detailed explanation of the YAML specification file, see the tutorial [How to specify an analysis with YAML](https://docs.immuneml.uio.no/tutorials/how_to_specify_an_analysis_with_yaml.html). + +See also the following tutorials for specific instructions: +- [Training ML models](https://docs.immuneml.uio.no/tutorials/how_to_train_and_assess_a_receptor_or_repertoire_classifier.html) for repertoire classification (e.g., disease prediction) or receptor sequence classification (e.g., antigen binding prediction). In immuneML, the performance of different machine learning (ML) settings can be compared by nested cross-validation. These ML settings consist of data preprocessing steps, encodings and ML models and their hyperparameters. +- [Exploratory analysis](https://docs.immuneml.uio.no/tutorials/how_to_perform_exploratory_analysis.html) of datasets by applying preprocessing and encoding, and plotting descriptive statistics without training ML models. +- [Simulating](https://docs.immuneml.uio.no/tutorials/how_to_simulate_antigen_signals_in_airr_datasets.html) immune events, such as disease states, into experimental or synthetic repertoire datasets. By implanting known immune signals into a given dataset, a ground truth benchmarking dataset is created. Such a dataset can be used to test the performance of ML settings under known conditions. +- [Applying trained ML models](https://docs.immuneml.uio.no/tutorials/how_to_apply_to_new_data.html) to new datasets with unknown class labels. +- And [other tutorials](https://docs.immuneml.uio.no/tutorials.html) + + +### Command line usage + +The `immune-ml` command takes only two parameters: the YAML specification file and a result path. +An example is given here: + +```bash +immune-ml path/to/specification.yaml result/folder/path/ +``` + +For each instruction specified in the YAML specification file, a subfolder is created in the +`result/folder/path`. Each subfolder will contain: +- An `index.html` file which shows an overview of the results produced by that instruction. Inspecting the results of an immuneML analysis typically starts here. +- A copy of the used YAML specification (`full_specification.yaml`) with all default parameters explicitly set. +- A folder containing all raw results produced by the instruction. +- A folder containing the imported dataset(s) in optimized binary (Pickle) format. + +## Support + +We will prioritize fixing important bugs, and try to answer any questions as soon as possible. We may implement suggested features and enhancements as time permits. + +If you run into problems when using immuneML, please see [the documentation](https://docs.immuneml.uio.no/latest/). In particular, we recommend you check out: +- The [Quickstart tutorial](https://docs.immuneml.uio.no/latest/quickstart.html) for new users +- The [Troubleshooting](https://docs.immuneml.uio.no/latest/troubleshooting.html) page + +If this does not answer your question, you can contact us via: +- Twitter [`@immuneml`](https://twitter.com/immuneml) +- Email [`contact@immuneml.uio.no`](mailto:contact@immuneml.uio.no) + +To report a potential bug or suggest new features, please [submit an issue on GitHub](https://github.com/uio-bmi/immuneML/issues). + +If you would like to make contributions, for example by adding a new ML method, encoding, report or preprocessing, please [see our developer documentation](https://docs.immuneml.uio.no/latest/developer_docs.html) and [submit a pull request](https://github.com/uio-bmi/compairr/pulls). + +## Requirements + +- [Python 3.7 or 3.8](https://www.python.org/) +- Python packages: + - [airr](https://pypi.org/project/airr/) (1 or higher) + - [dill](https://pypi.org/project/dill/) (0.3 or higher) + - [editdistance](https://pypi.org/project/editdistance/) (0.5.3 or higher) + - [fishersapi](https://pypi.org/project/fishersapi/) + - [gensim](https://pypi.org/project/gensim/) (3.8 or higher, < 4) + - [logomaker](https://pypi.org/project/logomaker/) (0.8 or higher) + - [matplotlib](https://matplotlib.org) (3.1 or higher) + - [matplotlib-venn](https://pypi.org/project/matplotlib-venn/) (0.11 or higher) + - [numpy](https://www.numpy.org/) (1.18 or higher, but at most 1.23.5) + - [pandas](https://pandas.pydata.org/) (1 or higher) + - [plotly](https://plotly.com/python/) (4 or higher) + - [pystache](https://pypi.org/project/pystache/) (0.5.4) + - [Pytorch](https://pytorch.org/) (1.5.1 or higher) + - [PyYAML](https://pyyaml.org) (5.3 or higher) + - [regex](https://pypi.org/project/regex/) + - [scikit-learn](https://scikit-learn.org/) (0.23 or higher) + - [scipy](https://www.scipy.org) + - [tensorboard](https://www.tensorflow.org/tensorboard) (1.14.0 or higher) + - [tzlocal](https://pypi.org/project/tzlocal/) +- Optional dependencies when using DeepRC: + - [DeepRC](https://github.com/ml-jku/DeepRC) (0.0.1) + - [widis-lstm-tools](https://github.com/widmi/widis-lstm-tools) (0.4) + - [tqdm](https://tqdm.github.io/) (0.24 or higher) + - [h5py](https://www.h5py.org/) (2.10.0 or lower when using DeepRC 0.0.1) +- Optional dependencies when using TCRdist: + - [parasail](https://pypi.org/project/parasail/) (1.2) + - [tcrdist3](https://github.com/kmayerb/tcrdist3) (0.1.6 or higher) + +# Citing immuneML + +If you are using immuneML in any published work, please cite: + +Pavlović, M., Scheffer, L., Motwani, K. et al. The immuneML ecosystem for machine learning analysis of adaptive immune +receptor repertoires. Nat Mach Intell 3, 936–944 (2021). https://doi.org/10.1038/s42256-021-00413-z + + + +<hr> + + +© Copyright 2021-2022, Milena Pavlovic, Lonneke Scheffer, Keshav Motwani, Victor Greiff, Geir Kjetil Sandve + + + + + + +%prep +%autosetup -n immuneML-2.2.5 + +%build +%py3_build + +%install +%py3_install +install -d -m755 %{buildroot}/%{_pkgdocdir} +if [ -d doc ]; then cp -arf doc %{buildroot}/%{_pkgdocdir}; fi +if [ -d docs ]; then cp -arf docs %{buildroot}/%{_pkgdocdir}; fi +if [ -d example ]; then cp -arf example %{buildroot}/%{_pkgdocdir}; fi +if [ -d examples ]; then cp -arf examples %{buildroot}/%{_pkgdocdir}; fi +pushd %{buildroot} +if [ -d usr/lib ]; then + find usr/lib -type f -printf "\"/%h/%f\"\n" >> filelist.lst +fi +if [ -d usr/lib64 ]; then + find usr/lib64 -type f -printf "\"/%h/%f\"\n" >> filelist.lst +fi +if [ -d usr/bin ]; then + find usr/bin -type f -printf "\"/%h/%f\"\n" >> filelist.lst +fi +if [ -d usr/sbin ]; then + find usr/sbin -type f -printf "\"/%h/%f\"\n" >> filelist.lst +fi +touch doclist.lst +if [ -d usr/share/man ]; then + find usr/share/man -type f -printf "\"/%h/%f.gz\"\n" >> doclist.lst +fi +popd +mv %{buildroot}/filelist.lst . +mv %{buildroot}/doclist.lst . + +%files -n python3-immuneML -f filelist.lst +%dir %{python3_sitelib}/* + +%files help -f doclist.lst +%{_docdir}/* + +%changelog +* Fri Jun 09 2023 Python_Bot <Python_Bot@openeuler.org> - 2.2.5-1 +- Package Spec generated @@ -0,0 +1 @@ +39081600c4bebf8532bc2eb916d182a1 immuneML-2.2.5.tar.gz |
